A 'One in a Million' Shot to Tackle Dopamine-Linked Brain Disorders
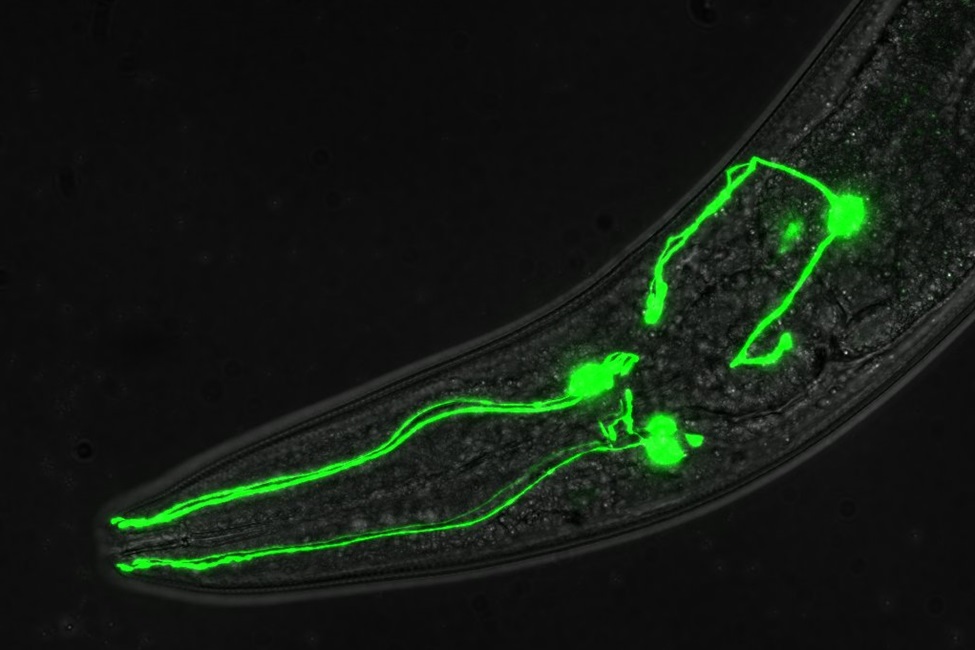
Image of C. elegans head showing dopamine neurons labeled with green fluorescent protein. (Photo credit: Peter Rodriguez, Jr., Florida Atlantic University)
Dopamine, a powerful brain chemical and neurotransmitter, is a key regulator of many important functions such as attention, experiencing pleasure and reward, and coordinating movement. The brain tightly regulates the production, release, inactivation and signaling of dopamine via a host of genes whose identity and link to human disease continue to expand.
Brain disorders associated with altered dopamine signaling include substance use disorder, attention deficit hyperactivity disorder (ADHD), autism, bipolar disorder, schizophrenia and Parkinson’s disease. The complexity of the human brain and its dopamine-associated disorders have encouraged many researchers to seek insights from simpler organisms whose genes bear striking similarity to those found in humans and where opportunities for genetic insights to disease can be pursued more efficiently and inexpensively.
With the help of a tiny, transparent worm called Caenorhabditis elegans, researchers from Florida Atlantic University have identified novel players in dopamine signaling by taking advantage of a powerful platform generated via the Million Mutation Project (MMP) for the rapid identification of mutant genes based on their functional impact.
The key resource of the MMP is a collection of 2,007 worm strains bearing chemically induced gene mutations. The genomes of each strain have been fully sequenced, the information archived and accessible via the web, and with all strains available for research use. Altogether, the MMP collection or “library” contains more than 800,000 unique genetic changes. On average, each gene in the worm's genome has about eight different mutations that change the protein produced, offering multiple opportunities to link gene disruption to changes in physiology and behavior.
“We turned to C. elegans to more efficiently elucidate the genetic, molecular and cellular bases of neural signaling than we could with rodent models,” said Randy D. Blakely, Ph.D., senior author, executive director of the FAU Stiles-Nicholson Brain Institute, the David J.S. Nicholson Distinguished Professor in Neuroscience, and a professor of biomedical science in FAU’s Schmidt College of Medicine. “It turns out that the proteins involved in dopamine regulation in C. elegans are highly conserved across evolution, suggesting that lessons learned from a simpler organism with a much simpler ‘brain’ could provide clues to dopamine linked disorders or how to better treat them.”
Nearly 20 years ago, Blakely’s team identified a profound behavioral change in worms when dopamine signaling is altered termed Swimming-induced-paralysis (Swip).
“We found that an inability to constrain the actions of dopamine leads worms to freeze in a few minutes when placed in water, whereas normal worms will thrash about for up to 60 minutes or more,” said Blakely.
To find new genes involved in dopamine signaling, Osama Refai, Ph.D., first author and a former research assistant professor; Peter Rodriguez, Jr., co-author and a graduate student; and Zayna Gichi, co-author and a research assistant, all within the Blakely Lab, tested 300 lines in the MMP library to find worms that showed Swip behavior. They used a dopamine signaling blocker to see if the worms could resume swimming, thereby verifying that excess dopamine signaling was the cause for their Swip. From that point, with that strain’s mutations already mapped to specific genes, the group could move very quickly to identify the gene whose change caused paralysis.
The results of this effort, reported in the Journal of Neurochemistry, included novel mutations in the worm gene encoding the dopamine transporter (dat-1), which functions to vacuum away dopamine from synapses after release, and that had previously been used to identify the Swip phenotype.
“Although, finding mutations in dat-1, a gene we already knew about didn’t accomplish our goal, this finding gave us confidence that our screen worked as intended, and that discoveries might lie ahead of us in the mutated genome of our other Swip lines,” said Blakely.
Indeed, further Swip screening revealed a surprising gene whose mutation produced Swip in worms and in humans, leads to a rare genetic disorder known as Bardet-Biedl Syndrome (BBS). Mutations in BBS arise in multiple proteins that together function as a larger protein complex termed the BBSome. Consistent with a larger protein complex at work, Blakely’s team found that mutations in all worm BBSome homologs produced Swip.
The complex of BBSome proteins is known to play a crucial role in the transport of proteins and lipids within the cell, and prominently into tiny hair-like extensions possessed by many cells termed primary cilia. The dopamine neurons of the worm possess primary cilia that allow the worm to sense by touch the world around them.
Over the past few years, scientists have found that many if not all neurons in the mammalian brain possess primary cilia that also can regulate cell signaling. According to Blakely, BBSome proteins are at work ensuring these protrusions carry the proper number and kinds of channels and receptors that will define capacities for cell signaling.
“Our results indicate that loss of BBS-1 in worm dopamine neurons results in excess signaling by the neurotransmitter, known to inhibit movement-controlling motor neurons. One mechanism we are considering involves a role of BBS-1 and other BBSome proteins in escorting dat-1 encoded protein to the cell surface to keep extracellular dopamine levels low and thereby not allow a completely shutting down of movement,” said Blakely. “Indeed, in an earlier screen, we identified another gene whose mutation acts exactly that way, and we found that overexpressing this gene in our BBS-1 mutant rescued full swimming behavior.”
In prior studies, Blakely and his team employed chemical mutagenesis to mutate the genomes of worms to find Swip mutants. Although, in these efforts, it took the group six months or more to identify the veritable “needle in the haystack,” a hunt for a single DNA base change among the millions making up the worm genome.
“Compared to our previous screening efforts, the MMP based approach allowed us a significant speed enhancement. Rather than map and sequence to identify the mutations in the strain, we could simply look up the known mutations in this line and then narrow down the culprit gene by testing specific candidates directly and almost immediately,” said Blakely.
Employing the MMP library, the researchers were able to screen more than 23,000 single nucleotide mutations across 300 MMP strains and nominate candidate genes within a few days after identifying a line showing dopamine-dependent Swip. The researchers’ initial behavioral screening effort covered about 15% of the MMP library and resulted in the identification of 10 strains, nine of which are now in the queue for new gene identification.
“Given the significant medical impact of altered dopamine signaling in multiple neurobehavioral disorders, further studies of how BBSome proteins regulate the dopamine transporter may lead to new strategies for treatment,” said Blakely.
-FAU-
Latest News Desk
- STEM Teachers in High-need Schools Resilient Despite ChallengesAn FAU College of Education researcher collaborated on a study examining 30 years of STEM teacher trends, focusing on qualifications and changes in high-need, primarily high-poverty U.S. schools.
- Alcohol-Related Deaths in the U.S. More than Double from 1999 to 2020FAU researchers explored overall trends as well as by age, gender, race and region. The sharpest spike occurred among 25-34-year-olds (nearly fourfold), while individuals aged 55-64 had the highest rates.
- Hyman Science Lecture Series at FAU Hosts Journalist Carl ZimmerThe Nat and Dorothy Hyman Science Lecture Series, as part of the Schmidt College of Science at FAU, will host science journalist Carl Zimmer on Thursday, Nov. 21 at 5:30 p.m.
- FAU REPORT: Consumers Face Elevated Prices Despite Waning InflationThe years of high inflation appear to be over as inflation is now in line with the Federal Reserve's target, though prices will likely remain permanently elevated, according to the Monthly Inflation Report.
- FAU Engineers Win NSF Grant for Unique School of AI Bio-Robotic FishA school of bio-robotic fish with advanced sensors will create a versatile network capable of maneuvers and station-keeping beyond current vehicles in shallow coastal areas and in deep ocean environments.
- FAU Tech Runway® Hosts 2024 Maverick ReceptionFlorida Atlantic University's Tech Runway® will host its 2024 Maverick Reception on Wednesday, Nov. 13 from 6 to 8 p.m. at its site, 901 NW 35th St., Boca Raton campus.